Research

Liu, Dali
Title/s: Professor
Graduate Program Director
Office #: FH-422
Phone: 773.508.3093
Email: dliu@luc.edu
Degrees
- 1988 B.S., Wuhan University, China
- 2001 Ph.D., University of Oklahoma
- 2002-2003 Postdoctoral Fellow, University of Oklahoma
- 2003-2004 Postdoctoral Fellow, UT Southwestern Medical Center at Dallas
- 2004-2009 Postdoctoral Fellow, Brandeis University
Research Interests

Dr. Liu’s research group primarily employs X-ray crystallography in combination with mechanistic, biological and computational methods to study proteins involved in bacterial infections and other human diseases. Following the doctrine of “Structure Determines Function”, our research evolves around determining key protein structures that are mechanistically informative on biochemical processes. Putting together the obtained structures like snapshots, key knowledge on the biological functions of those critical proteins can be deduced. We aim to apply this knowledge to discover novel antimicrobial methods and conduct structure-based drug design.
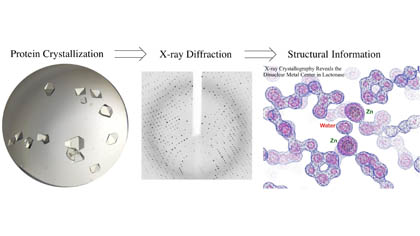
Current Projects:
1. Discover Novel Antimicrobial Methods to Combat Antibiotic Resistance.
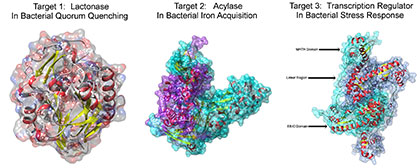
We currently focus on three protein targets involved in difference aspects of virulence expression in pathogenic bacteria. Target 1: on-going studies aim to employ lactonase as a protein therapeutic to disrupt bacterial cell-to-cell communication, quorum-quenching. On Target 2, on-going studies aim to inhibit acylase to disrupt bacterial iron acquisition. On Target 3, on-going studies aim to modulate GabR-dependent transcription regulation to weaken bacterial stress response. These targets are critical for different biological functions but converge on virulence expression in persistent infections such as the lung infection in patients with cystic fibrosis. The targeted pathogens include Pseudomonas aeruginosa, Burkholderia cepacia complex, Klebsiella pneumonia et al, all of which are known for their ability to resist widely used antibiotics such as penicillin. Overall, a variety of approaches including protein engineering, inhibitor design and discovery of super agonistic ligands will be used to exploit the potential of reducing virulence, to bring about novel treatments that overcome antibiotic resistance in bacteria, and to improve the quality of life of patients who suffer from these life-long infections.
2. Structural Enzymology and Structure Based Drug Design
Through collaborative efforts, we are also interested in elucidating the mechanism of allosteric regulators and rational drug design against protein targets in humans. The enzymes in these studies include bacterial Sucrose Synthase and bacterial ADP-Glucose Pyrophosphorylase, which are important in biofuel production. Mammalian GABA Aminotransferase and human Ornithine Aminotransferase are studied as drug targets in neurological disorder and hepatic cancer respectively.
Selected Publications
Current publications via PubMed
1. NCBI Bibliography https://www.ncbi.nlm.nih.gov/myncbi/1T9O9viZidtkZ/bibliography/public/
2. Google Scholar https://scholar.google.com/citations?user=1drmkSwAAAAJ&hl=en
3. Loop https://loop.frontiersin.org/people/281796/publications
